Summary
Professor James Tour of Rice University presents a novel molecular robot that was developed thanks to crucial experimental work with Robert Pal of Durham University that targets a specific cell, and then drills into and through the cell wall causing the cell guts to spill out. Preliminary targets are either a cancerous cell or an antibiotic-resistant bacterial cell. The mechanism is rapid cell death via mechanical tearing, not a chemical mechanism, that is highly resistant to both cancer mutations and also antibiotic-resistant bacteria mutations. Neighboring cells are left unaffected.
While this technology is available in vitro now, Professor James Tour expects this technology to be clinically proven and also available to no-option end-oflife patients within 1-5 years, with full approval available in 5-15 years.
Presenters

James Tour, Rice University
James M. Tour, a synthetic organic chemist, is at the Center for Nanoscale Science and Technology at Rice University in 1999 where he is presently the T. T. and W. F. Chao Professor of Chemistry, Professor of Computer Science, and Professor of Materials Science and NanoEngineering. In 2020, he became a Fellow of the Royal Society of Chemistry…

Nicolas Giuseppone, University of Strasbourg
Nicolas Giuseppone received his PhD in asymmetric catalysis (laboratory of Prof. H.B. Kagan, Orsay University), performed a post-doctoral research in total synthesis (laboratory of Prof. K.C. Nicolaou, The Scripps Research Institute, La Jolla, CA, USA), and entered the field of supramolecular chemistry…
Presentation: THE 4TH MODALITY OF MEDICAL TREATMENT: CELLULAR SURGERY
PROJECTIONS
Potential applications achievable in the near term (1-5 years) and the challenges preventing them now.
• Clinical proof, translation, and no-option patient use is achievable in 1-5 years Challenges
• Hiring of synthetic chemists – we need the world to open back up after COVID
• Fundraising for Nanorobotics LLC was slowed due to COVID-restricted travel
• Suitable scale-up operations for synthesis
• Final selection of the specific target molecular motor
Potential applications achievable in the long term (~30 years) and their
associated challenges.
• This 4th modality of medical treatment should be a mainstay approach
- Prior work from the Tour lab demonstrates the world’s fastest nano car.
- The Tour lab Molecular Drive Nano Car is capable of not only high speeds with recorded 6,000 RPM but also capable of driving around targeted pylons and passing through the two end goal posts.
- The Molecular Nanomechanical Car is driven by light and heat, providing energy to the freespinning double bond. This provides unidirectional spinning of the wheels and forward motion across the gold surface.
- The Molecular Nanomachine Drill operates with some similar principles to the car. The differences are the use of cell targeting and targeted cell adherence. The drill utilizes a similar light activation power source as the car to spin the rotor.
These holes depicted are open for at least one minute. The holes are big enough and last long enough that the inner machinery of the target cell will spill out of the cell, causing a necrotic death. The cells do not die of apoptosis via a chemical method, they die of a physical mechanical action and immediate necrosis.
- The Fluorophore is what adheres to the cell wall. The targeting method and the adherence method can be adjusted. If the drill is not adhered to the cell wall before activating the light, the drill will simply spin free and not drill into a cell.
- The light activation energizes the freely rotating double bond between the stator and rotor. The molecular drill now has the enduring power while the light is on to be able to drill into the cell wall. If the drill is adhered but the double bond is not energized with light, the rotor can swing and spin around via the weak molecular forces present between the cell wall and the stator, but the rotor will not have the energy needed to drill into the cell.
- The Molecular drill has different attachments to target different cells.
- Propidium Iodide fluoresces when it is in contact with DNA. This is how we know that the cell was penetrated not only past the first cell wall but also penetrating the nucleus of the cell via a mechanical action.
- Some cell lines in the studies are damaged by light, and we prove in other slides that the light activation alone is not responsible for the dramatic necrotic death of target cells.
- The effects of nanomachines 3 and 4, and control molecule 5 on PC-3 cells upon UV- activation; the rate of necrotic cell death and permeabilization of analytes into the cells was recorded. The UVexposure times are shown in each image.
- (a) Blank cells without molecular motors
- (b) with introduction of 3
- (c) introduction of 4
- (d) with non-active 5
- (e) Identical imaging sequence using 3 with the introduction of 100 nM PI confirming molechanical cell permeabilization with intercalation of RNA and DNA. All scale bars = 20 μm.
- Individual cells can be targeted and killed by the Molecular Nano Drill Robot, leaving a direct neighbor cell unaffected. This is a 3D schematic showing the x and z planes above the glass cover slip, and also the top-down x and y plane view of the two cells. The PC3-targeted fast motor version robot #4 was able to target, adhere, and mechanically drill into the target cell when activation was performed with 2 photons at 710 nm wavelength near infrared. The direct neighbor cell contacting the target cell was left unaffected.
- Demonstrating precise selectivity in cell death using 2- photon 710 nm NIR light and targeted nanomachines for MCF7 and PC-3 cell lines.
- Cell lines studied are effected by UV light causing ROS damage. This shows that the molecular drills are causing the damage and subsequent necrotic cell death, and that the molecular drills are actually quenching ROS oxidation.
- Clonogenic Assay: Long-Long-Term survival at 9 days.
- Tumors are obliterated by the Molecular Nano Drill [MNM + Light]with one treatment.
- New bacteria strains are continually emerging that are completely resistant to all antibiotics that we have available to kill them. By 2050 10 million people per year could be dying by antibiotic resistant bacteria alone.
- Meanwhile, there is a thirty-five year gap with absolutely no new antibiotics that could treat these new antibiotic resistant bacteria strains.
- Molecular Drills kill the antibiotic tolerant, persister cells.
- Reduction in abundance (expressed as log N/N0) of persister cells following treatment with standard antibiotics and 2x MIC of different visible light activated MNMs or 1% DMSO followed by irradiation with 291 mW/cm2 of 405 nm light.
- Molecular Drills disrupt the biofilms and reduce the biomass.
- Biofilms are recalcitrant to antibiotic treatment due to multiple tolerance mechanisms (phenotypic resistance). This causes persistence of biofilm infections in spite of antibiotic exposure which predisposes to antibiotic resistance development (genetic resistance). Reduction in biofilm biomass of P. aeruginosa and S. aureus following 15, 30, and 45 min of irradiation (291 mW cm-2, 405 nm light) in the presence of 2x MIC of antibiotic, 1% DMSO or 2x MIC of MNMs DL-654, DL-877, and DL-878 in the presence and absence of light. Biofilm biomass was determined using the crystal violet method. Each bar represents the mean ± SEM of at least 3 biological replicas. Asterisks denote the statistical significance level of the comparison between irradiated samples and the corresponding dark controls determined using a Kruskal- Wallis test in GraphPadPrism8.*=p<0.05, **=p<0.01
- https://pubmed.ncbi.nlm.nih.gov/28407419/
- Repeated exposure to Molecular Drills does not cause resistance in the bacteria culture. Repeated exposure to existing antibiotics does cause resistance to the treatment.
- TEM photographs of E. coli following exposure to 405 nm light in the presence of 1% DMSO or 0.5x MIC of different visible light activated MNMs. Arrows denote different types of ultrastructural changes observed in MNM-treated samples compared to DMSO-treated ones. Note the leakage of cellular organelles denoted by the blue arrow.
Presentation: Molecular machines that work at all scales
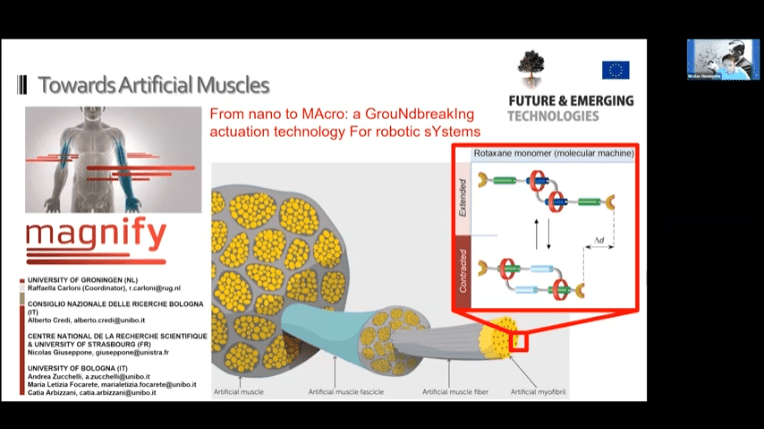
- Hierarchical mechanics of human muscle – multiple molecular machines work together to create work
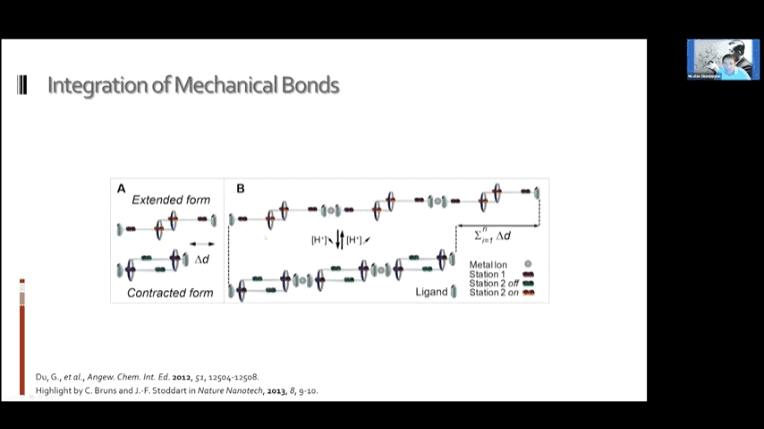
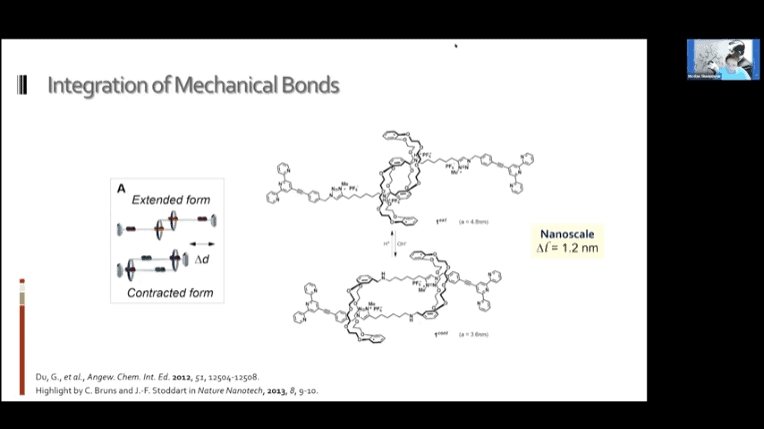
- Molecular sliding units mimicking the human muscle. These monomers can be daisy-chained to create a nanoscale muscle mechanism.
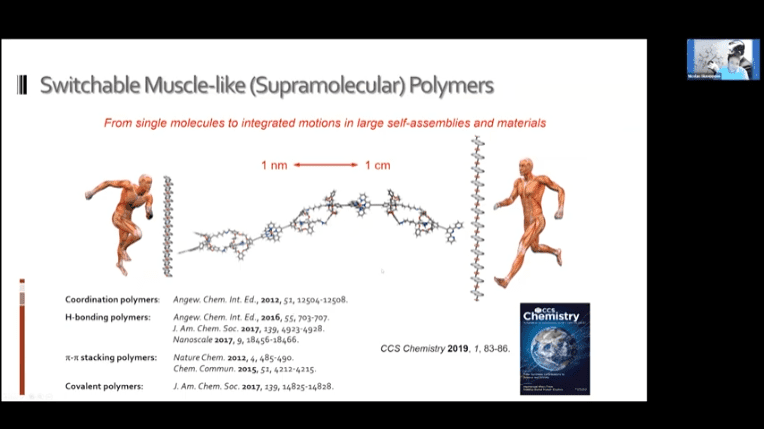
- Switchable polymer muscles can be built that approach the utility of real human muscles
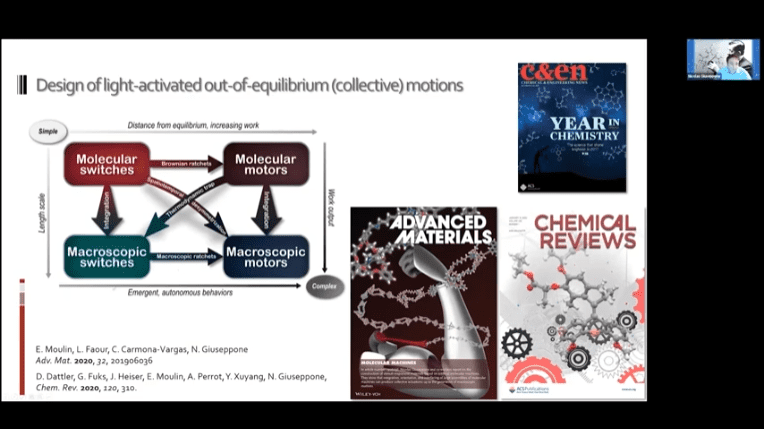
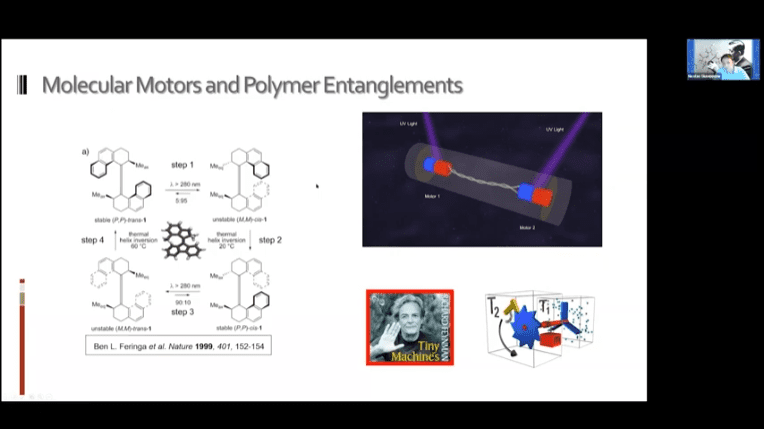
- A new mechanism of nanoscale muscle – molecular motors twist fibers around each other to create contraction.
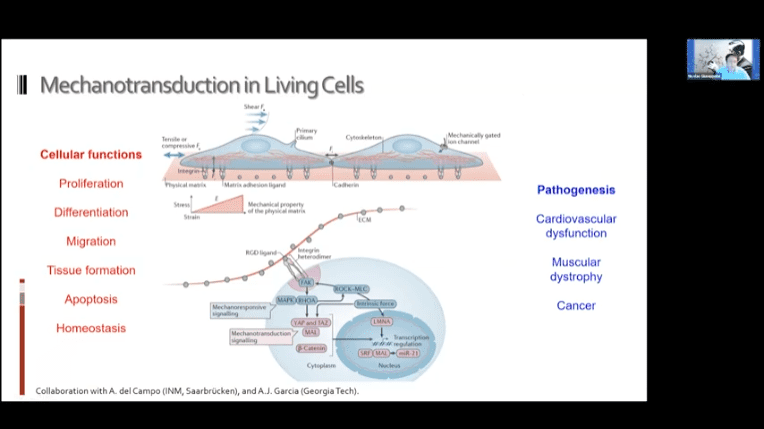
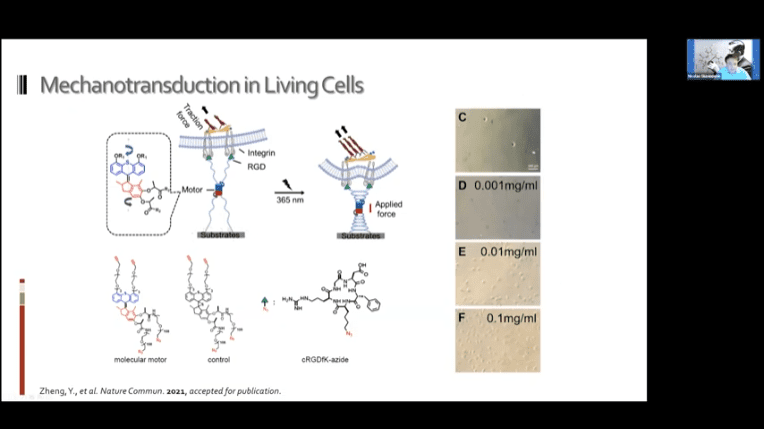
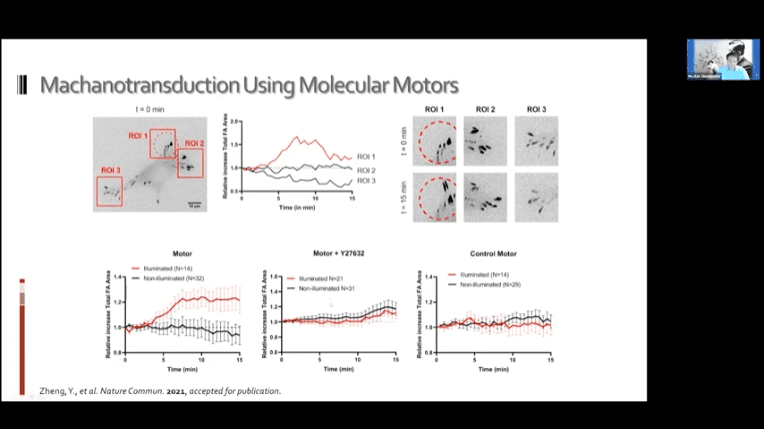
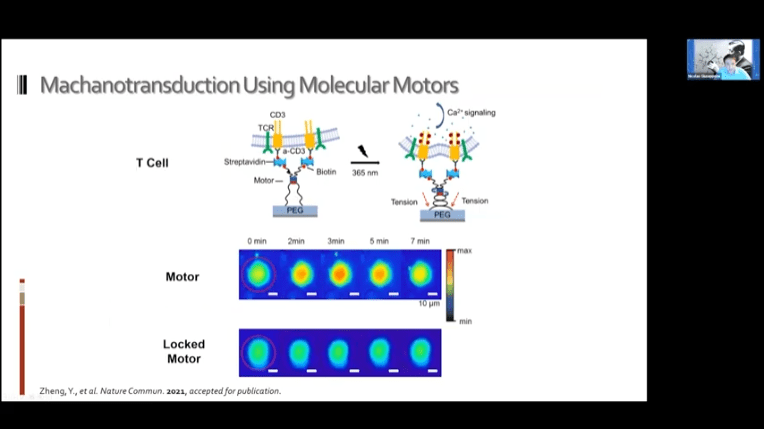
- It may be possible to use these systems to achieve mechanotransduction in living cells. The idea is to bind the motors to the cell in two places and then to either a ligand or other target substrate and drag the two objects together.
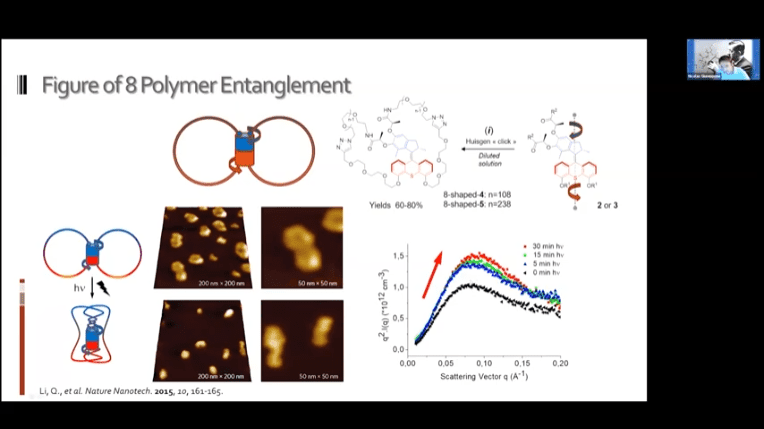
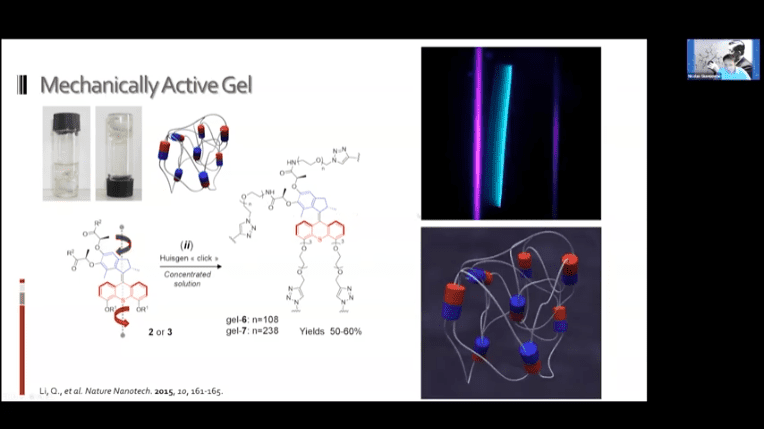
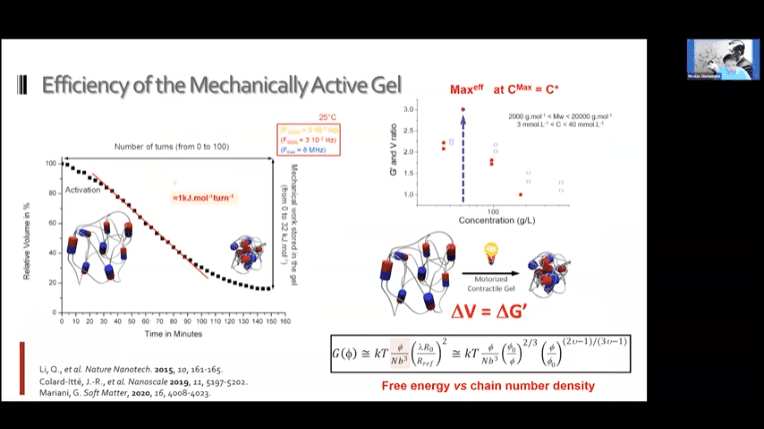
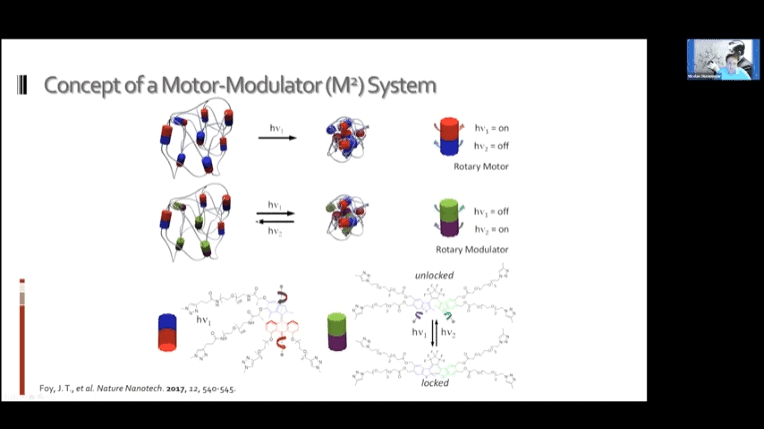
- Creating a loose network of these motors connected to each other by polymer threads leads to a mechanically active gel – a large mass of molecular machines that can contract and expand on demand.
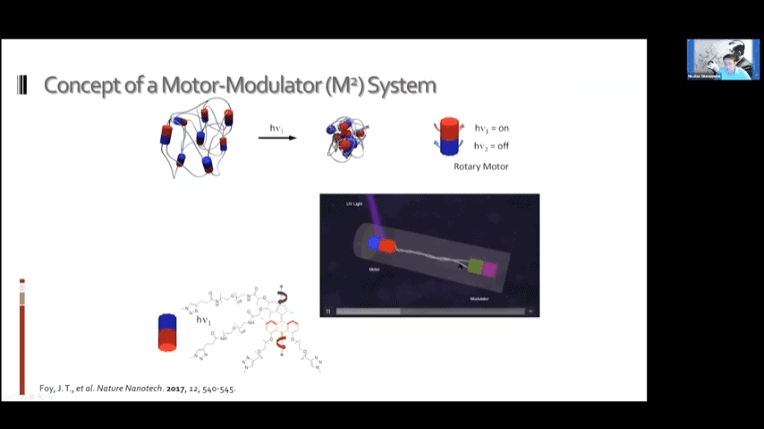
The fundamental concept is the same as the mechanotransduction model, but modulators can be added to reverse the contraction and return the gel to its original state.
- Incorporating light switches into these motors may lead to fast, strong, effective synthetic artificial muscle.
- Short, medium, and long term challenges for nanoscale artificial muscles
Acknowledgments
Seminar summary by Tim Potter.