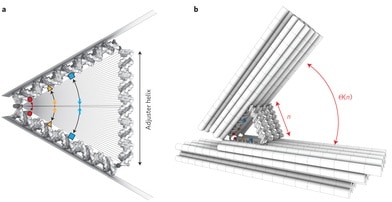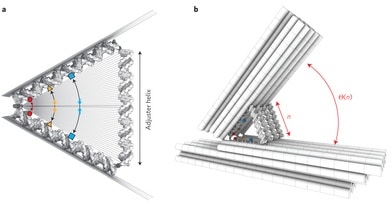
The powerful and elegant molecular recognition code that makes possible double helical DNA has also made scaffolded DNA origami and its parent field, structural DNA nanotechnology, probably the most widespread and useful approach to bottom-up molecular nanotechnology. The 2-nm diameter of the DNA helix leads to the suspicion, however, that the ultimate precision obtainable with these technologies might not be much better than 5 nm, more than an order of magnitude less precise than atomic precision. A paper published two months ago, however, demonstrates much finer control using a DNA hinge with “adjuster helices” to control the angle of the hinge. The abstract (reference numbers omitted) from “Placing molecules with Bohr radius resolution using DNA origami“:
Molecular self-assembly with nucleic acids can be used to fabricate discrete objects with defined sizes and arbitrary shapes … . It relies on building blocks that are commensurate to those of biological macromolecular machines and should therefore be capable of delivering the atomic-scale placement accuracy known today only from natural and designed proteins …. However, research in the field has predominantly focused on producing increasingly large and complex, but more coarsely defined, objects … and placing them in an orderly manner on solid substrates … . So far, few objects afford a design accuracy better than 5 nm … , and the subnanometre scale has been reached only within the unit cells of designed DNA crystals … . Here, we report a molecular positioning device made from a hinged DNA origami object in which the angle between the two structural units can be controlled with adjuster helices. To test the positioning capabilities of the device, we used photophysical and crosslinking assays that report the coordinate of interest directly with atomic resolution. Using this combination of placement and analysis, we rationally adjusted the average distance between fluorescent molecules and reactive groups from 1.5 to 9 nm in 123 discrete displacement steps. The smallest displacement step possible was 0.04 nm, which is slightly less than the Bohr radius. The fluctuation amplitudes in the distance coordinate were also small (±0.5 nm), and within a factor of two to three of the amplitudes found in protein structures … .
The Bohr radius is the most probable distance between the proton and the electron of a hydrogen atom in its ground state, and is approximately 52.9 pm (0.0529 nm). This work from Hendrik Dietz’s group at the Technical University of Munich, Germany, continues his work on dynamic nanomachines from DNA nanotechnology that we cited last spring. For additional information about the new paper, see:
A hat tip to Next Big Future for this description pointing to two more detailed descriptions. “Bootstrapping Atomic Precision using adjustable DNA origami triangle hinge“:
A new study demonstrates that researchers can control the distance between two molecules such that they can adjust the step size to as small as Bohr’s radius. This proof-of-concept study using DNA origami techniques shows how molecular positioning can be fine-tuned at the atomic level at room temperature in solution. This work has applications for molecular architecture as well as templated chemical reactions …
From Alex Klotz at Physics Forums Insights “Atomic Positioning with DNA Hinges“:
… The authors created a hinge out of DNA which they selectively open or close by varying the length of one of the DNA molecules, which allows them to adjust the positions of molecules attached to the hinge by as little 0.04 nanometers, much less than the typical spacing between atoms in a solid. …
… By adjusting it one base-pair at a time, from 10 to 50 base-pairs, roughly 3 to 17 nanometers (a good rule of thumb is that double stranded DNA has three base-pairs per nanometer), they adjusted in opening angle of the hinge in 41 discrete steps. Looking at the hinges with a transmission electron microscope, it is clear that the angle can be precisely controlled. …
From Heather Zeiger at Phys.org “Using a DNA scaffold to place molecules with Bohr’s radius resolution“:
… This proof-of-concept study using DNA origami techniques shows how molecular positioning can be fine-tuned at the atomic level at room temperature in solution. This work has applications for molecular architecture as well as templated chemical reactions. …
Zeiger describes how Funke and Dietz used TEM (transmission electron microscopy) images to determine how the angle of the hinge changes with the length of the adjuster helices, how they used FRET (Förster resonance energy transfer) to measure the effects of hinge angle on the interaction of chromophores attached to the hinge components, and how they used chemical crosslinking reactions to determine how thermal fluctuations at room temperature affect molecular distances.
This proof-of-concept study demonstrates the possibility of using a DNA scaffold to control molecular distance. When we asked about the implications of his research, Dr. Funke said, “Arranging matter with ever more precision is a key goal for science and technology. Our study shows, that scaffolded DNA origami enables the rational positioning of two molecules with atomic resolution and therefore opens up new opportunities to study and manipulate molecular interactions.”
Almost a year ago we cited work by another research group (at the Ohio State University) demonstrating structural DNA nanotechnology with programmed motions, including a tunable hinge joint. This work a continent away further reveals an emerging network of researchers pushing the limits of what can be done with DNA nanotechnology. Will next year see atomically precise positioning leading to positioning atoms or molecular fragments to achieve atomically precise positional control of chemical synthesis?
—James Lewis, PhD