A select set of videos from the 2013 Foresight Technical Conference: Illuminating Atomic Precision, held January 11-13, 2013 in Palo Alto, have been made available on vimeo. Videos have been posted of those presentations for which the speakers have consented. Other presentations contained confidential information and will not be posted.
The fourth speaker at the Commercial Scale Devices – Part 2 session, the winner of the 2012 Feynman Prize for Theoretical work, David Soloveichik, presented his prize-winning work “Artificial Biochemistry with DNA” https://vimeo.com/62119584 – video length 29:14. Dr. Soloveichik began his talk by asking if we could recapitulate the feats of biology, specifically computation with networks of molecular interactions, with de novo engineering. After the basic technology is developed, possible applications could include artificial control modules that could be inserted into cells to create “smart drugs”, or as control modules for completely artificial systems (“wet robots”). Dr. Soloveichik has made the slides from his talk available here.
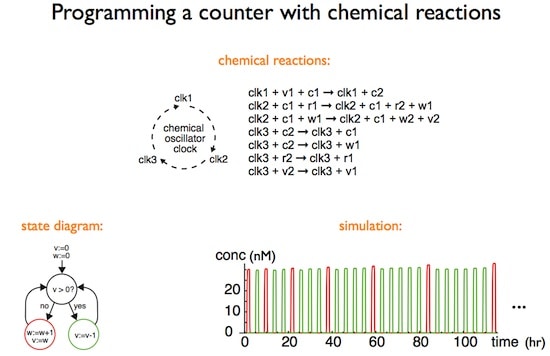
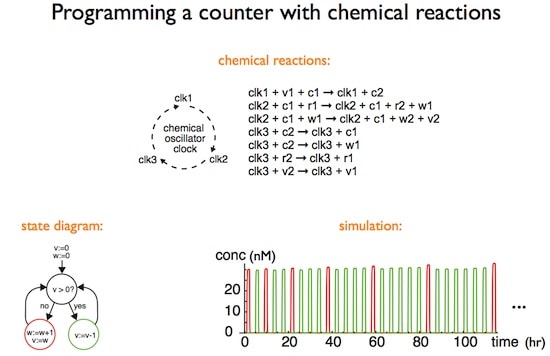
Dr. Soloveichik noted that living cells are controlled by regulatory networks that are complex not only because of their size, but also because of their dynamic properties and kinetic behavior. As an example, he cited the cyanobacterial circadian clock oscillator, in which limit cycle oscillations can continue for 10 days in vitro due to mutual binding and phosphorylation-dephosphorylation reactions. In these control networks, the chemical interactions are for the purpose of processing information so that network structure is more important than the chemical identity of the interacting species.
To create a technology from scratch capable of producing complex molecular systems, perhaps exceeding the results of biology, Dr. Soloveichik pointed to the need for a material whose interactions can be programmed. Nucleic acids are an ideal choice because to make two single strands bind to each other, it is only necessary to make one the Watson-Crick complement of the other. 3-D structure and sequence are not important; only what parts are complementary and can bind other parts is important. He calls sequences that effectively act as units “domains” and gives them numbers, indicating W-C complement by a star (*). Two identical strands competing for the same binding partner can cause strand displacement, or branch migration, which occurs readily without the help of enzymes. Bernie Yurke was the first to realize that strand displacement can be used to rationally engineer molecular scale changes in structure. What was missing was a flexible strategy to chain or cascade such changes together to create functional networks.
The system, Dr. Soloveichik explained, obeys three rules obeyed in all possible ways: the bind rule, the release rule, and the displace rule. There are two kinds of domains: long and short. The short domain is short enough that its binding is readily reversible. The bind rule says that two single strand domains that are W-C complements of each other can bind. THe release rule says that any strand bound by a short domain can unbind. The key displace rule says that you can displace a domain by an identical domain, but only if an adjacent domain is already bound. the physical basis of this rule is that there is a large kinetic barrier to beginning strand displacement, but co0localization can overcome it by increasing the effective local concentration. The rate of displacements can be controlled over six orders of magnitude by changing the length or the binding strength of the short domain. Dr. Soloveichik illustrated these rules with an AND logic gate constructed from a small number of domains. Ne noted a 2011 Science paper (“Scaling Up Digital Circuit Computation with DNA Strand Displacement Cascades” by Qian and Winfree) in which the square-root of a 4-bit number was calculated by a circuit that comprised 130 DNA strands. He also mentioned Microsoft’s Visual DNA Strand Displacement formal language (DSD) for describing and modeling strand displacement cascades, and a flash game developed by David Baker’s group
To demonstrate what strand displacement cascades can do, Dr. Soloveichik presented a theoretical argument that strand displacement cascades can in principle mimic the temporal dynamics of any other chemical system, implying they can exhibit any complex computational or regulatory function possible in a chemical system. He illustrated this claim with the strand displacement implementation of a bimolecular reaction X + Y –> Z at rate k. X, Y, and Z correspond to certain DNA strands; production and consumption correspond to these strands being free or bound, etc. Several examples were presented.
—James Lewis, PhD