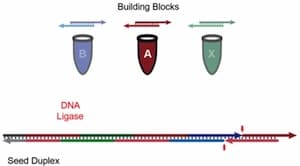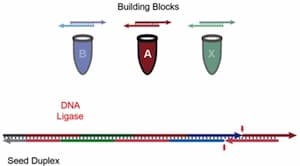
A few months ago we cited a cheaper, easier way developed by researchers at McGill University to build long DNA scaffolds. A further substantial improvement, in which long DNA segments of up to 1000 base pairs are produced less expensively than the short 100-base strands they previously used is described at KurzweilAI.net “A new technique to build complex custom-designed DNA scaffolds“:
McGill University researchers have devised a new technique to produce long, custom-designed DNA strands to build nanoscale structures to deliver drugs to targets within the body or take electronic miniaturization to a new level.
Researchers have been assembling and experimenting with DNA structures or “DNA origami” for years, as KurzweilAI has reported. But as these applications continue to develop, they require increasingly large and complex strands of DNA. It can take hundreds of these short strands to assemble nanotubes for applications such as smart drug-delivery systems.
That poses a problem: automated systems used for making synthetic DNA can’t produce strands containing more than about 100 bases (the chemicals that link up to form the DNA strands). …
Additional details are provided in a McGill University news release “A better way to build DNA scaffolds“:
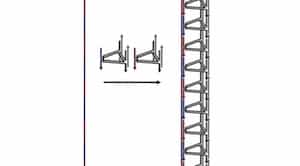
… In new research published May 5 in Nature Communications [abstract], however, Sleiman’’s team at McGill reports that it has devised a technique to create much longer strands of DNA, including custom-designed sequence patterns. What’s more, this approach also produces large amounts of these longer strands in just a few hours, making the process potentially more economical and commercially viable than existing techniques.
The new method involves piecing together small strands one after the other, so that they attach into a longer DNA strand with the help of an enzyme known as ligase. A second enzyme, polymerase, is then used to generate many copies of the long DNA strand, yielding larger volumes of the material. The polymerase process has the added advantage of correcting any errors that may have been introduced into the sequence, amplifying only the correctly sequenced, full-length product. …
The team used these strands as a scaffold to make DNA nanotubes, demonstrating that the technique allows the length and functions of the tubes to be precisely programmed. “In the end, what we get is a long, synthetic DNA strand with exactly the sequence of bases that we want, and with exactly as many repeat units as we want,” explains Sleiman, who co-authored the study with Graham Hamblin, who recently completed his doctorate, and PhD student Janane Rahbani.
“This work opens the door toward a new design strategy in DNA nanotechnology,” Sleiman says. “This could provide access to designer DNA materials that are economical and can compete with cheaper, but less versatile technologies. In the future, uses could range from customized gene and protein synthesis, to applications in nanoelectronics, nano-optics, and medicine, including diagnosis and therapy.”
As the research paper’s abstract points out:
Generally, only a small portion of a DNA structure needs to be addressable, while the rest is purely structural. Scaffolds with specifically placed unique sites in a repeating motif greatly minimize the number of components used, while maintaining addressability.
By providing a way to build complex objects with very few building blocks, this work points toward structural DNA nanotechnology that is less expensive, more scalable, and more commercially applicable.
—James Lewis, PhD