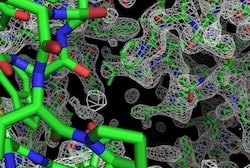
With the development of artificial molecular machines still at an early stage, natural biological molecular machines, mostly protein molecules, still provide most information about how molecular machines work. Crucial to extracting this information is knowledge of the 3D structures of these molecules, usually obtained by arduous analysis of X-ray diffraction of protein crystals. Scientists in the US and UK have now reported software advances that will allow many more molecular machines to be studied. A hat tip to Phys.Org for reprinting this Los Alamos National Laboratory news release “Mysteries of ‘molecular machines’ revealed“:
Phenix software uses X-ray diffraction spots to produce 3-D image
Scientists are making it easier for pharmaceutical companies and researchers to see the detailed inner workings of molecular machines.
“Inside each cell in our bodies and inside every bacterium and virus are tiny but complex protein molecules that synthesize chemicals, replicate genetic material, turn each other on and off, and transport chemicals across cell membranes,” said Tom Terwilliger, a Los Alamos National Laboratory scientist. “Understanding how all these machines work is the key to developing new therapeutics, for treating genetic disorders, and for developing new ways to make useful materials.”
To understand how a machine works you have to be able to see how it is put together and how all its parts fit together. This is where the Los Alamos scientists come in. These molecular machines are very small: a million of them placed side by side would take up less than an inch of space. Researchers can see them however, using x-rays, crystals and computers. Researchers produce billions of copies of a protein machine, dissolve them in water, and grow crystals of the protein, like growing sugar crystals except that the machines are larger than a sugar molecule.
Then they shine a beam of X-rays at a crystal and measure the brightness of each of the thousands of diffracted X-ray spots that are produced. Then researchers use the powerful Phenix software, developed by scientists at Los Alamos, Lawrence Berkeley National Laboratory, Duke and Cambridge universities, to analyze the diffraction spots and produce a three-dimensional picture of a single protein machine. This picture tells the researchers exactly how the protein machine is put together.
The 3-D advance
Recently Los Alamos scientists worked with their colleagues at LBNL and Cambridge University to make it even easier to visualize a molecular machine. In a report in the journal Nature Methods [abstract] this month, Los Alamos scientists and their team show that they can obtain three-dimensional pictures of molecular machines using X-ray diffraction spots that could not previously be analyzed.
Some molecular machines contain a few metal atoms or other atoms that diffract X-rays differently than the carbon, oxygen, nitrogen, and hydrogen atoms that make up most of the atoms in a protein. The Phenix software finds those metal atoms first, and then uses their locations to find all the other atoms. For most molecular machines, however, metal atoms have to be incorporated into the machine artificially to make this all work.
The major new development to which Los Alamos scientists have contributed was showing that powerful statistical methods could be applied to find metal atoms even if they do not scatter X-rays very differently than all the other atoms. Even metal atoms such as sulfur that are naturally part of almost all proteins can be found and used to generate a three-dimensional picture of a protein. Now that it will often be possible to see a three-dimensional picture of a protein without artificially incorporating metal atoms into them, many more molecular machines can be studied.
Cracking the cascade
Molecular machines that have recently been seen in three-dimensional detail include a “huge” molecular machine called Cascade that was reported in the journal Science [abstract] this summer. The Cascade machine is present in bacteria and can recognize DNA that comes from viruses that infect the bacteria. The Cascade machine is made up of 11 proteins and an RNA molecule and looks like a seahorse, with the RNA molecule winding through the whole “body” of the seahorse. If a foreign piece of DNA in the bacterial cell is complementary to part of the RNA molecule then another specialized machine can come by and chop up the foreign DNA, saving the bacterium from infection.
Los Alamos and Cambridge University scientists who were developing the Phenix software were part of the team that visualized this protein machine for the first time. The Phenix software has been used to determine the three-dimensional shapes of over 15,000 different protein machines and has been cited by over 5000 scientific publications.
About Phenix
Phenix is a collaborative team developing new software for determining the shapes of molecular machines using X-ray crystallography. Phenix includes researchers from Lawrence Berkeley National Laboratory, Los Alamos National Laboratory, Duke University, and Cambridge University. …
The Phenix suite of Python programs and more information can be obtained at the Phenix web site. As usual with advances in basic technical capabilities, it is difficult to predict the consequences for either near-term pharmaceutical research or long-term understanding of how complex molecular machine systems function. The eventual development of high throughput atomically precise manufacturing, otherwise termed molecular manufacturing, will surely require scientific insights and engineering innovations beyond what can be gleaned from the study of biological molecular machines, just as the development of modern aircraft required insights beyond those derived from studying birds. Nevertheless, at this early stage, such insights will surely prove useful.
—James Lewis, PhD