The earliest proposal to “open a path to the fabrication of devices to complex atomic specifications” envisioned designing proteins to fold in predetermined ways. Over the years we have cited here numerous advances along this path, most recently here, here, here, here, and here. There has also been interest in polymers that are chemical cousins of proteins, for example, the peptoids, in which the side chains are appended to the nitrogen atom in the peptide bond joining the monomers, rather than to the alpha carbon atom, as is the case with proteins. Four years ago we cited a report of initial successes in the rational design of peptoids, speculated that it might be opening a path to advanced nanotechnology, but noted that their successful computer modeling had so far only produced structures composed of nine subunits. Now we can cite further work by a subset of that group reporting a peptoid nanosheet, two molecules thick but extending laterally for micrometers, based on a structural element not seen in the natural world. A hat tip to Nanowerk for reprinting this news release from the U.S. Dept.of Energy Office of Science “Understanding and Predicting Self-Assembly“:
To mimic complex natural proteins’ capabilities as sensors, catalysts, and more using synthetic materials that can withstand industrial conditions, scientists must first know how to finesse the building blocks they’ll use. Molecular Foundry staff worked with scientists to discover a new design rule that controls the way in which polymer building blocks adjoin to form the backbones that run the length of tiny biomimetic sheets.
Understanding the rules that govern how the nanosheets self-assemble could be used to piece together complex sheet structures and others, such as nanotubes and crystalline solids, that could be customized into incredibly sensitive chemical detectors or long-lasting catalysts, to name a few possibilities.
Scientists aspire to build nanostructures that mimic the complexity and function of nature’s proteins, but are made of durable and synthetic materials. These microscopic widgets could be customized into incredibly sensitive chemical detectors or long-lasting catalysts, to name a few possible applications.
But as with any craft that requires extreme precision, researchers must first learn how to finesse the materials they’ll use to build these structures. A recent discovery by scientists at the Molecular Foundry is a big step in this direction.
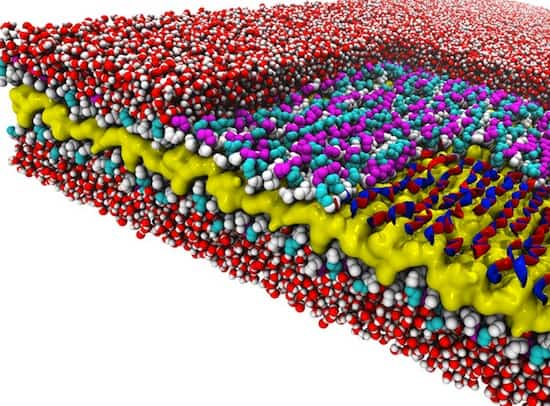
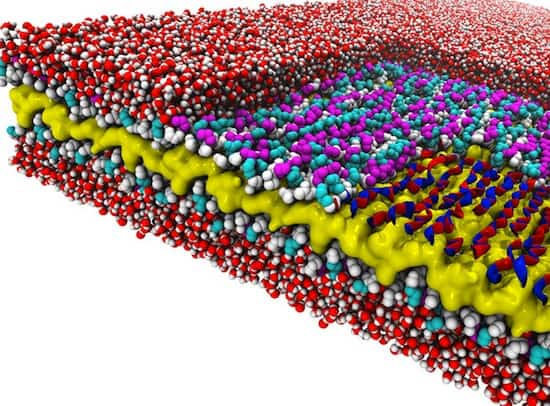
The researchers discovered a design rule that enables a recently created material to exist. The material is a peptoid nanosheet. It’s a flat structure only two molecules thick, and it’s composed of peptoids, which are synthetic polymers closely related to protein-forming peptides.
The design rule controls the way in which polymers adjoin to form the backbones that run the length of nanosheets. Surprisingly, these molecules link together in a counter-rotating pattern not seen in nature. This pattern allows the backbones to remain linear and untwisted, a trait that makes peptoid nanosheets larger and flatter than any biological structure.
The Molecular Foundry scientists say this never-before-seen design rule could be used to piece together complex nanosheet structures and other peptoid assemblies such as nanotubes and crystalline solids.
What’s more, they discovered it by combining computer simulations at National Energy Research Scientific Computing Center, another DOE user facility located at Berkeley Lab, with x-ray scattering and imaging methods to determine, for the first time, the atomic-resolution structure of peptoid nanosheets.
The research was reported in a paper published last fall in Nature [abstract]. Additional details are contained in a longer release from the Lawrence Berkeley Lab “Newly discovered ‘design rule’ brings nature-inspired nanostructures one step closer“:
“This research suggests new ways to design biomimetic structures,” says Steve Whitelam, a co-corresponding author of the Nature paper. “We can begin thinking about using design principles other than those nature offers.”
Whitelam is a staff scientist in the Theory Facility at the Molecular Foundry, a DOE Office of Science user facility located at Berkeley Lab. He led the research with co-corresponding author Ranjan Mannige, a postdoctoral researcher at the Molecular Foundry; and Ron Zuckermann, who directs the Molecular Foundry’s Biological Nanostructures Facility. They used the high-performance computing resources of the National Energy Research Scientific Computing Center (NERSC), another DOE Office of Science user facility located at Berkeley Lab.
Peptoid nanosheets were discovered by Zuckermann’s group five years ago. They found that under the right conditions, peptoids self assemble into two-dimensional assemblies that can grow hundreds of microns across. This “molecular paper” has become a hot prospect as a protein-mimicking platform for molecular design.
To learn more about this potential building material, the scientists set out to learn its atom-resolution structure. This involved feedback between experiment and theory. Microscopy and scattering data gathered at the Molecular Foundry and the Advanced Light Source, also a DOE Office of Science user facility located at Berkeley Lab, were compared with molecular dynamics simulations conducted at NERSC.
The research revealed several new things about peptoid nanosheets. Their molecular makeup varies throughout their structure, they can be formed only from peptoids of a certain minimum length, they contain water pockets, and they are potentially porous when it comes to water and ions.
These insights are intriguing on their own, but when the scientists examined the structure of the nanosheets’ backbone, they were surprised to see a design rule not found in the field of protein structural biology.
Here’s the difference: In nature, proteins are composed of beta sheets and alpha helices. These fundamental building blocks are themselves composed of backbones, and the polymers that make up these backbones are all joined together using the same rule. Each adjacent polymer rotates incrementally in the same direction, so that a twist runs along the backbone.
This rule doesn’t apply to peptoid nanosheets. Along their backbones, adjacent monomer units rotate in opposite directions. These counter-rotations cancel each other out, resulting in a linear and untwisted backbone. This enables backbones to be tiled in two dimensions and extended into large sheets that are flatter than anything nature can produce.
“It was a big surprise to find the design rule that makes peptoid nanosheets possible has eluded the field of biology until now,” says Mannige. “This rule could perhaps be used to build many more unrealized structures.”
Adds Zuckermann, “We also expect there are other design principles waiting to be discovered, which could lead to even more biomimetic nanostructures.”
Perhaps the most striking result from this work is the discovery that protein-like polymers can adopt secondary structures entirely unlike anything seen with natural proteins. The zigzag pattern that enables the formation of the nanosheet is termed by the authors a Σ (sigma) strand. The authors explain that the Σ strand results from the ability of adjacent monomers to rotate in opposite directions, while most elements of protein secondary structure are built from one rotational state. The authors note that this difference “might greatly expand the repertoire of folded polymeric building blocks.” They conclude taht this new building principle combined with a wide selection of chemically diverse monomers “offers a way to create new structured polymers through combinatorial design.”
—James Lewis, PhD
Discuss these news stories on Foresight’s Facebook page or on our Facebook group.