We found 164 results for your search.
One way to reach molecular machine systems is to get really, really good at protein engineering. If that’s your goal, you’ll want to be in Boston on May 17-21 for PEGS 2010, “the essential protein engineering summit”. Not sure if this is your pathway? Just reading the talk titles is educational. And they have great… Continue reading The protein engineering path to molecular manufacturing
Presenter Julian Englert, Adaptyv Bio Julian Englert co-founded Adaptyv Bio in 2020 and currently holds the role of Co-Founder. Before that, they worked at Xsensio as an R&D Engineer from September 2018 to October 2019. Julian Englert completed their Bachelor of Science in Materials Science from the Karlsruhe Institute of Technology (KIT) from 2012 to… Continue reading Julian Englert | Making a Protein Printer That Turns Bits Into Molecules @ MSD Workshop 2023
Presenter Simon Dürr, Ecole Polytechnique Federale der Lausanne (EPFL) Simon Dürr is a PhD student at Ecole Polytechnique Federale der Lausanne (EPFL) in Switzerland. He works on methods to improve protein engineering of metalloproteins using deep learning and molecular modeling. He is also known for his work improving accessibility of machine learning models via easy… Continue reading Simon Dürr | Designing stable Metalloproteins using Deep Learning
Summary David Baker from the University of Washington presents breakthrough advancements in de novo protein design. Deep learning pattern recognition hallucinates the desired protein structure and also generates the correct peptide sequence for accurate folding, and predicted proteins are highly transferable to actual proteins produced in a lab. Results are easily transferable to the production… Continue reading Protein-based Assemblies and Molecular Machines | David Baker, University of Washington
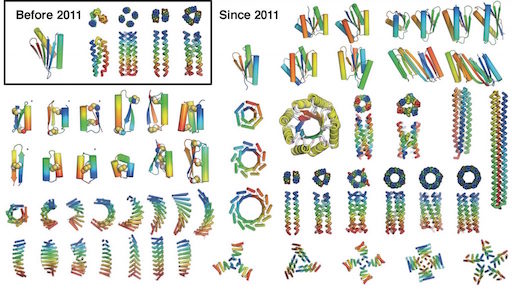
A review from the group leading recent rapid progress in de novo protein design describes the successes, identifies the remaining challenges, and heralds the advance “from the Stone Age to the Iron Age” in protein design.
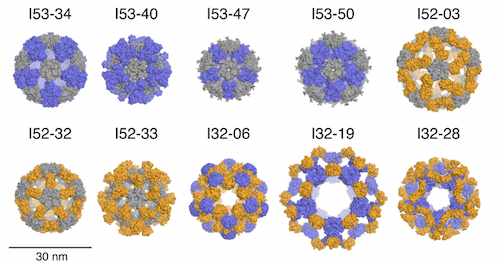
Ten designs spanning three types of icosahedral architectures produce atomically precise multi-megadalton protein cages to deliver biological cargo or serve as scaffolds for organizing various molecular functions.
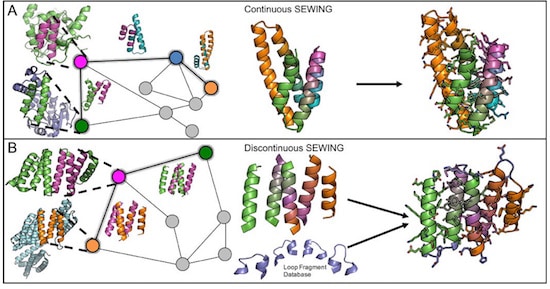
Computational recombination of small elements of structure from known protein structures generates a vast library of designs that balance protein stability with the potential for new functions and novel interactions.
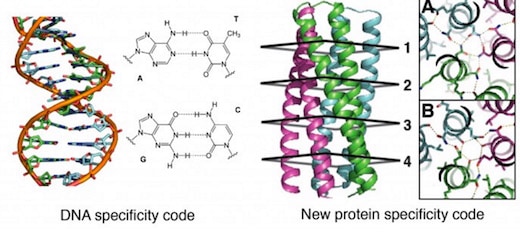
Computer designed networks of hydrogen bonds allow programming specific interactions of protein interfaces, facilitating programming molecular recognition.
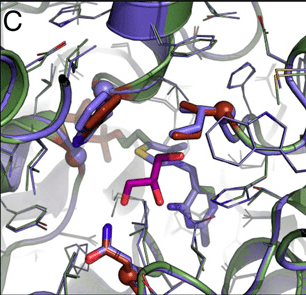
Computational design of an enzyme that carboligates three one-carbon molecules to form one three-carbon molecule, an activity that does not exist in nature, provides proof-of-principle for a novel metabolic pathway for carbon fixation.