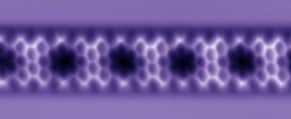
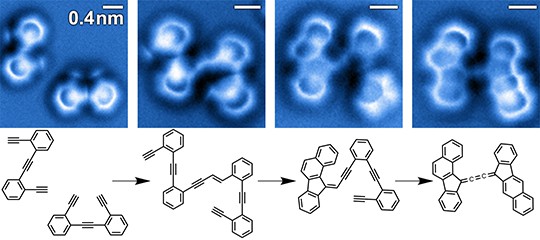
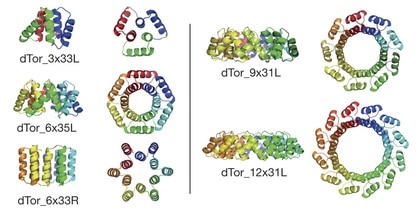
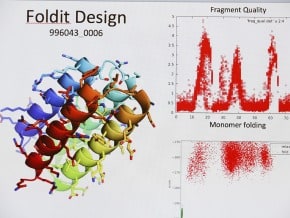
Combining computational nanotechnology with a noncontact-atomic force microscope probe tipped by a single CO molecule allowed researchers to visualize the dance of individual chemical bonds during a complex organic reaction on a silver surface.
Watching individual chemical bonds during a reaction